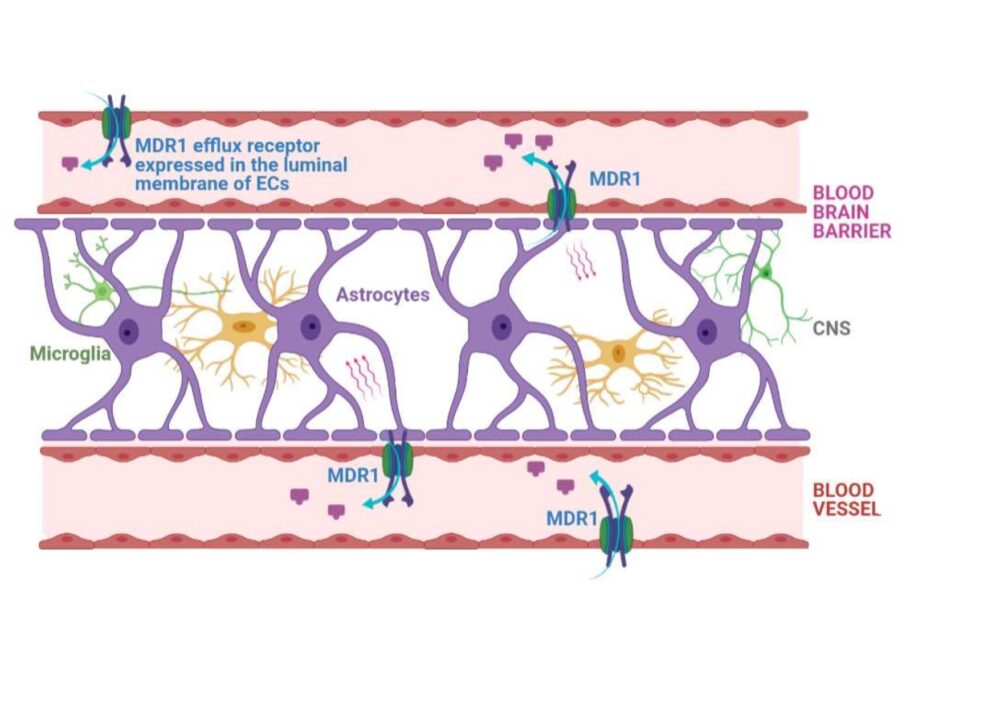
The pharmacological implications in blood-brain barrier permeability assays.
Drug delivery to the brain is a challenging evaluation to consider during novel compounds discovery stages to study central nervous pharmacokinetics in terms of efficacy and safety.
This is the fact to the specialized cellular integration of the blood-brain barrier (BBB) and their interaction with additional perivascular elements form a highly selective barrier that regulates substance movement in and out the Central Nervous System (CNS).
In the case of large molecules, movement across the BBB is dependent of carrier mediated transporters such as the multidrug resistance (MDR1) gene product, P-glycoprotein (Pgp) which according to the scientific community and main regulatory agencies, is considered the reference efflux transporter at the CNS barrier site, due to its pharmacological relevance.
In this context, it is, of tremendous importance to ascertain whether new molecular entities (NMEs) are potential substrates of P-gp in order to accurately predict their pharmacokinetics in the CNS at the early stages of drug discovery.
Following FDA recommendations, ReadyCell scientists have developed a cell-based model system to reliably predict P-pg-mediated drug interactions in the CNS. In addition, this cellular model which consists of Madin-Darby Canine Kidney (MDCKII) cells overexpressing the MDR1 gene product (P-pg) when combined with a patented Shipping Medium is able to be transported worldwide in a ready-to-use format.
Based on the scientific literature, we highlight the permeability values (Papp) of a data set of 7 marketed CNS drugs tested in the MDCKII-MDR1 cell model system.
| MARKETED CNS DRUGS | PRIMARY TRANSPORT MECHANISM | VALUES PAPP X 106 (CM/S) |
|---|---|---|
| Alprenolol | Transcellular | 53 +/- 7 |
| Atenolol | Paracellular | 0,7 +/- 0.2 |
| Baclofen | Paracellular/Influx | 0,7 +/-0,4 |
| Diazepam | Transcellular | 65 +/- 18 |
| Metronidazole | Transcellular | 11 |
| Midazolam | Transcellular | 42 +/- 6 |
| Propanolol | Transcellular | 38-50 |
In case you are developing NMEs with the mechanism of action based on CNS, we advise you to consult our product portfolio specially designed for and contact our specialists at reagents@readycell.com or through our form.